Some people, like Imelda Marcos and our new Dr. Isis, have a thing for fancy shoes.
I go crazy for gadgets.
technorati tags: iphone, DNA, molecules, molecular structure, molecular modeling, Science education
For my birthday this year, my family bought me a new iPhone! Yeah!
So, I've been killing several hours today filling it with cute little iPhone apps. Who knew one little phone could be so much fun?
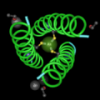
One app, I enjoy, is called Molecules.
Molecules lets you download structure files from the Protein Data Bank (PDB) and play with the structures on your phone!

Spreading your fingers makes the molecule larger.

Working with your fingers also lets you turn the molecule around.

It's also nice that the developer, Brad Larson, made the code available, 'cause I have all kinds of requests for improvements and new features.
Here's my wish list:
- Make the colors match the colors in Cn3D. Phosphorus should be green, not orange.
- In Cn3D, I can add annotations, I would like to be able to view those annotations.
- I would like to be able to see superimposed structures.
- I would like to be able to limit the view to selected parts of the structure.
- I would like to be able to change rendering and coloring styles.
- Add support for mmdb formatted files. I would be fine with making annotated files in Cn3D on my desktop computer and then viewing them in Molecules.
This sort of thing must be why I hear stories about schools getting iPhones for students.
That's so wonderfully geeky. I love it! :)
(Now all I need is an iPhone...)
Do you have to download the structures for this, or can you import your own PDB files?
Oh, and FYI, I think some colours are a discipline thing. In most branches of chemistry, phosphorus tends to be purple. Green is normally reserved for chlorine. ;)
I tried using both the PDB-formatted structures that I downloaded from the Protein Data Bank and PDB-formatted structures that I downloaded from the NCBI.
Only the structures from the Protein Data Bank worked.
I see your point about the colors, some of the viewers at the Protein Data Bank show phosphorus as orange, too.
I'm glad that you find it useful. If you're interested in my vision for the program, I was invited to write an article for the RCSB Protein Data Bank newsletter's educational column: http://www.rcsb.org/pdb/general_information/news_publications/newslette… .
I agree that there's a lot left to be done before the full educational and scientific potential of the application can be realized. I'll add mmdb as a file format for importing structures. I had hoped to get the ability to select NMR structures and overlay multiple structures in the latest update, but couldn't fit it in. I didn't spend as much time with metadata, so I'm not sure if the PDB format supports annotation, but I'll look at adding support in a way that won't clutter the limited display real estate of the portable device.
As far as coloring goes, I've used Jmol in the past, so I relied on its color scheme: http://jmol.sourceforge.net/jscolors/ . If there's a standard you could point me to, it would be simple to adjust the colors to match. Maybe a pop-up legend might help.
In regards to changing rendering styles, you can access a couple of simple ones via double-tapping on the display. Cartoon or ribbon visualization modes are high on my list of things to work on, and others have volunteered to help with that.
Unfortunately, I'm a little distracted right now with another educational / science application, so progress may be slow on this for the next couple of months.
This may be one of the hottest things I have ever seen in my life. I wonder if I could get an anatomy atlas for my iPhone? That would make surgery a whole lot easier when I am suddenly like, "Now wait, where is the spleen supposed to be again?"
You're in luck, Isis, check out this movie on YouTube.
Brad: Thanks for stopping by!
I haven't been able to find any documentation for the color scheme in Cn3D, but I will keep looking.
I'll definitely take a look at your article. I'm definitely interested. I don't know if it's helpful or not, but I do know that Cn3D is open source, so maybe that will be helpful.
Hey, I'm just glad to hear from researchers / educators that like the application. It makes me feel like this was worth it (a reassurance I sometimes need after reading some of the iTunes reviews).
You also mentioned that you were having trouble with non-RCSB PDB files. If you could point them out here or in the forums ( http://www.sunsetlakesoftware.com/forum ), it would help me to figure out what I'm doing wrong. I only tested the custom import functionality on a small number of non-RCSB files.
Thanks Brad,
I will post some examples in the forum.
ANIMATION DNA